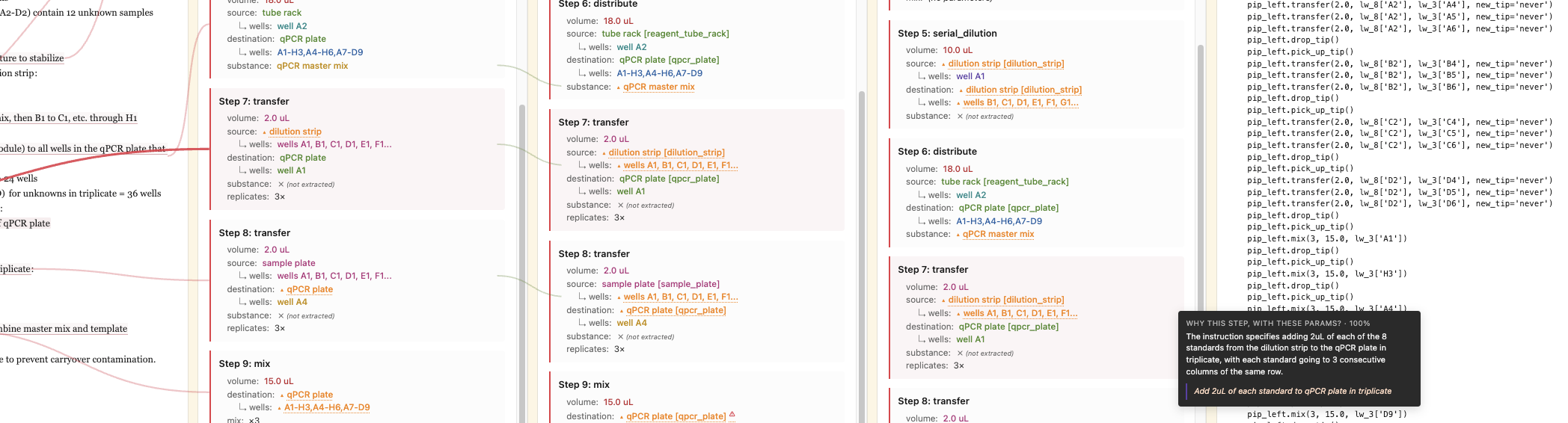
Turn natural-language lab protocols into Opentrons scripts — with citations for every extracted value.
Hover the marked phrases. Every value in the spec carries a citation like that — that’s the point.
- from instruction
- domain default
- inferred
- reviewer agreed
How it works
-
01 Extract
Sonnet reads your instruction and emits a structured spec. Every extracted value carries a claim: this came from the instruction, here’s the verbatim quote, or this was inferred, here’s the reasoning.
-
02 Resolve gaps
Deterministic suggesters fill what they can (config lookups, well-capacity defaults). Haiku audits non-deterministic suggestions against your instruction. Anything the system can’t decide goes to you, one decision at a time.
-
03 Simulate
The generated Python script runs through Opentrons’ own simulator before output. If the simulator rejects the script, we surface the failure loudly — no silent broken protocols.
What’s different
-
Every value carries provenance.
Other tools generate code and hope. Here, every extracted volume, well, labware, and substance comes with either the verbatim quote from your instruction or explicit reasoning if it was inferred. You can audit any decision the system makes.
-
An independent reviewer audits the LLM.
A smaller, separate model verifies every non-cited claim against your instruction. If it disagrees, you see the objection in the UI before deciding — not after the protocol fails on the robot.
-
The script is simulator-verified.
Opentrons’ own simulator runs the generated Python before output. Code that wouldn’t actually load on the robot never reaches you.